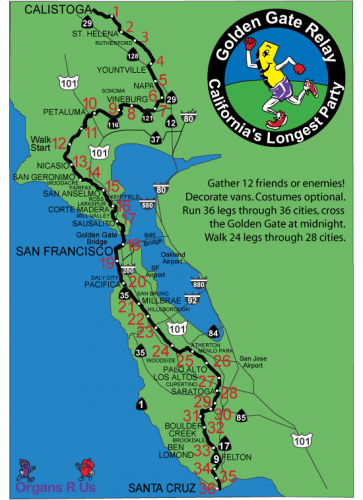
It’s been a wonderful, incredibly busy summer! After my classes ended in early May – and before starting summer research – I went out to California to visit family and to run in The Golden Gate Relay. Seeing family is always fantastic and the relay was a truly epic adventure. The relay course goes along the California coastline for nearly 200 miles, and it took our team about 24 hours to complete. Along the way (well, mostly before the race) we raised money for Organs ’R’ Us, a non-profit that promotes the need for organ donors and offers support for transplant recipients. The run was long and intense, but also a phenomenal experience – amazing teammates in the van and some transplant recipients at the finish line on the beach in Santa Cruz! And, yes, for some reason we’ve named our team Toxic Megacolon.
Once back in Ann Arbor, I moved to my new home and spent quite a bit of time and energy fixing everything up. I love so much about the place, but especially the view of the water from my bedroom window and the deer who visits daily (and gives me guilty stares when I catch him nibbling on my garden).
I’m over at the School of Public Health for the summer doing a research rotation in the Biostatistics Department and I’m absolutely loving it! Long-term, for my PhD and beyond, I am interested in working on statistical genetics projects, so I will rotate with a couple faculty that are part of the Center for Statistical Genetics at Michigan. There is a wide range of investigations currently underway here; part of why the summer has been such a good experience is that I’ve had the chance to learn more about the work of others in the group. It’s exciting how many directions the research could go, and working with people who do slightly-different-but-related research will help me narrow my own interests.
The past couple months, I’ve been working on a project where we’re getting ready to launch an app through social media that will end up (hopefully!) collecting thousands of health history surveys and (when people consent) saliva samples for genotyping. Because genetic effects are generally so small, enormous sample sizes are required for association studies. If we can learn to *predict* disease by identifying genetic risks, we can start with earlier intervention for patients. Essentially, in genetic association studies, we look directly at the link between genes and diseases, but biologically there are a few steps in between. It’s something along the lines of:
Gene –> transcript –> protein –> metabolite –> disease/phenotype
I’m excited about the project I’ve gotten involved with this summer, and it’s been great to be back on the research side of things. It’s also my first time really computing in a Unix environment, writing command line code, and figuring out how to navigate various buffers and shells in the data cluster. One of the main reasons I want to go into biostatistics is that it seems we’re at a point in biomedicine where we’re generating data faster than we’re learning how to use it. Particularly given the rapidly-evolving (no pun intended) genetic sequencing technology, I’m optimistic that there is much that remains to be discovered.
Finally, and completely unrelatedly, my friends and I celebrated the Summer Solstice with an evening party where we all wore colors found in sunsets! For the most part, it was simply another excuse to get together for a fun, summery night.